This post is somewhat of a follow-up to something that I wrote earlier this week. In bioinformatics, we often want to analyze all genes from an organism (or from multiple organisms). In many well-annotated genome databases, there is often a choice of isoforms available for each protein-coding gene, and the number of isoforms only ever seems to increase.
For example, in the latest set of human gene annotations (Ensembl 78), there are 406 protein-coding genes that have more than 25 transcripts. At one extreme, the human GPR56 gene has 77 transcripts, 61 of which are annotated as protein-coding! The length of these 61 putative protein products ranges from just 6 amino acids (!) all the way up to 693.
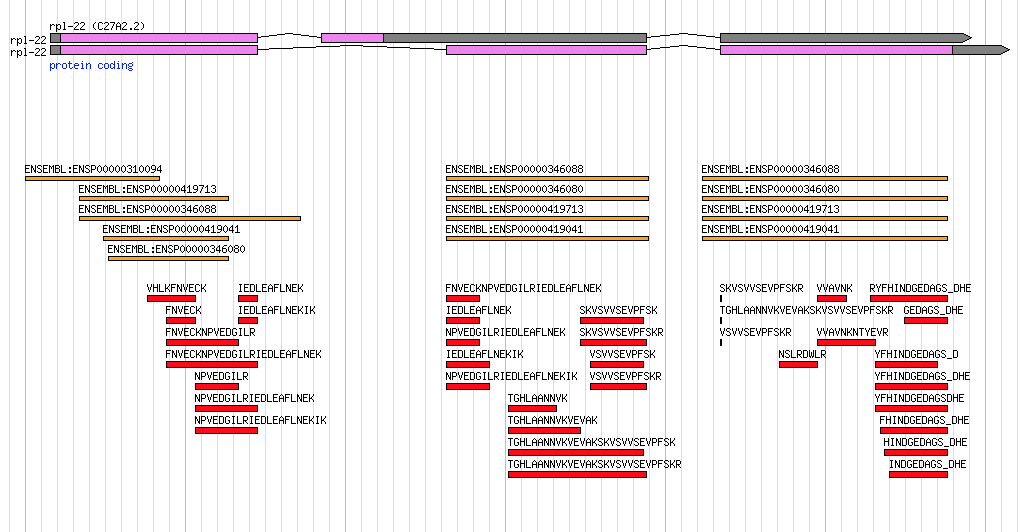
In Caenorhabditis elegans, sequence identifiers for genes were historically based on appending numbers to the identifier of the BAC/YAC/Cosmid clone containing that gene. E.g. B0348.1 would represent the first predicted gene on the B0348 clone, B0348.2 the second gene…and so on. When splice variants were discovered, curators appended letters for each isoform. E.g. B0348.2a and B0348.2b represent the two alternative isoforms of this gene. In the latest WS248 release of WormBase, one gene (egl-8) has 25 isoforms (all the way up to B0348.4y). I wonder what WormBase will do when a 27th isoform is discovered?
So how does one attempt to choose a single variant for use in a bioinformatics pipeline, and is this something that we should even be attempting? Historically, people have often opted for a quick-and-easy approach in order to get around this problem. Some examples from papers indexed by Google Scholar:
"In cases of alternative splicing, we chose the longest protein to represent a gene"
"In cases of multiple transcript isoforms, we chose the isoform with the longest CDS supported by transcript and protein homology in other mammalian species"
"Because of the redundancy of protein sequences, we chose only the longest isoform for every entry"
"In cases where a gene possesses more than one reference sequence, we chose the longest"
"When multiple protein entries are found for the same EntrezGene identifier, choose the longest sequence isoform"
This methodology is obviously not without problems (as others have reported on). So I'm genuinely curious as to what people do in order to choose a 'representative' isoform (whatever that means). The problem is further complicated when the reality might be that some genes consistently use different isoforms in different tissues or at different developmental time points.
Please comment below if you think you have found a good solution to this problem!